Sample introduction or the GC inlet parameters are very often overlooked when developing a GC or GC-MS method, but these parameters are so powerful that they can result in success or total failure of an application and as a result are also the main area for troubleshooting and maintenance within the GC system.
Obtaining the best resolution for the method is critical for accurate qualitative and quantitative analysis, especially where chromatographic resolution is relied upon with standard GC detectors, as there is no MSD to take advantage of analytical resolution.
Would you believe that only a small percentage of GC and GC-MS instruments are used to their full capabilities? This is the same for the data analysis systems.
In GC-MS, library integrity has massively evolved compared to 25 years ago. In those times the NBS library had 72000 compounds, this was superseded by a Wiley library of 230000 compounds in the late 90’s that for the first time contained multiple synonyms, greatly helping identifications. NIST libraries became mainstream around the year 2002 and are re-issued every three years.
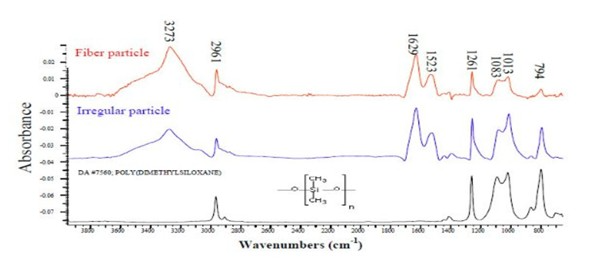
In FTIR – Hundreds of thousands of reference spectra are available via many suppliers but are often collated and offered on single websites. Full spectral searching, multiple regions searching, browsable libraries and batch searching for multiple samples are all widely available.
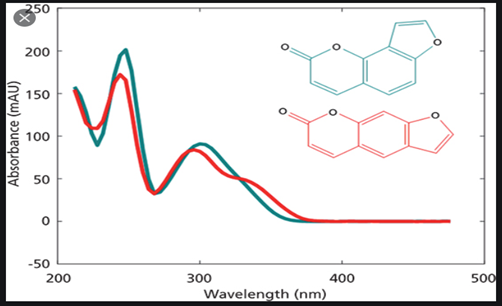
In LC – DAD the ability to collect UV-Vis 'scan' data enables peak purity estimations, this combined with a retention time identifier gives an overall 3-dimensional output akin to GC-MS.
In GC-MS, imagine if you suddenly didn't have access to the plethora of excellent Mass Spectral libraries. After the initial shock has subsided, thought processes should quickly lead to the fundamentals of Mass Spectral Interpretation (MSI)!
You'll be pleased to know there are many excellent tools available within this huge but fun area.
Imagine you have a sample extract with an unknown concentration of target analyte in a matrix you know has been difficult to extract from (leading to signal suppression) or which might conversely, have artificially enhanced the target response.
You don't have the time for a full method validation, so what do you do?
Techniques such as mass spectrometry can produce 10's or 100's of thousands of data points per sample. In these cases, it is impossible to satisfy the statistical experiments' design goal of having substantially more sample replicates than variable. If you have lots of extracted compound peaks, then some will appear to be significantly different purely by chance.
What can we do to help this?
A look at the differences between GC and LC sample introduction.
© 2005–2026 Anthias Consulting Ltd – Experts in Analytical Chemistry Consulting and Training.